Imagine a job could change your life.
Patients faced with life-changing illnesses find a path to healing at Mayo Clinic. We humanize the practice of health care and inspire hope in the people who need it most—one patient at a time. Our team-focused approach brings leading expertise to each patient, with research and education that deliver innovation. That's life changing.
Effective June 5, 2023, Mayo Clinic will no longer mandate COVID-19 vaccination as a condition of employment. Due to system limitations, our application may continue to reference COVID-19 vaccination as a requirement, for a period of time. If you have an offer or start date on or after June 5, 2023, you will not be required to be vaccinated.
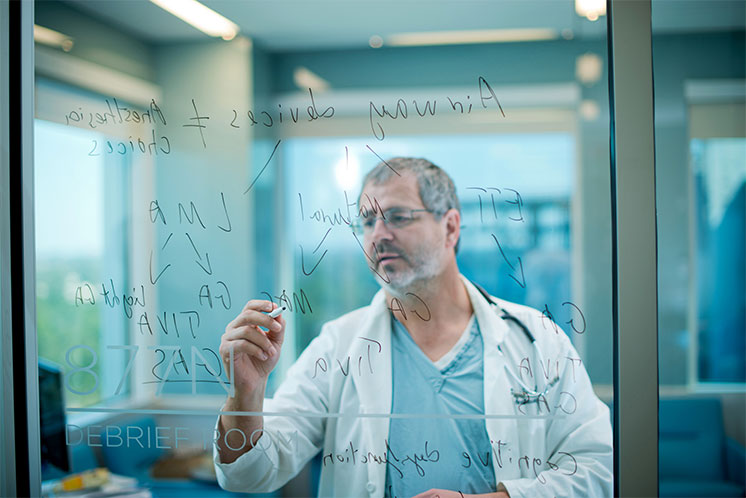

Physicians
At Mayo Clinic, you are a colleague of some of the most talented, experienced physicians in the world. You work with patients, conditions, and cases that most doctors will never encounter in their professional lives. In our physician-led environment, you will discover a culture of teamwork, professionalism and mutual respect where the needs of the patient always come first.
View open positions
Nursing
Explore limitless nursing opportunities at Mayo Clinic. From seasoned RNs to new grads, our tailored programs empower you to shape your career. Join a culture of excellence and make a difference in healthcare. Discover your path with Mayo Clinic Nursing today!
View open positions
Health Professionals
Mayo Clinic has a legacy of inspiring hope and contributing to health and well-being by providing the best care to every patient through integrated clinical practice, education and research. As a health professional, you will be part of an amazing team committed to solving the most serious and complex medical challenges–one patient at a time.
View open positions
Non-Medical
Envision yourself working for a global leader in an industry fueled by innovation and growth. You’ll find a world of non-medical focused careers with the power to change lives. Whether you’re a new graduate or an experienced business professional, have advanced degrees or a high school diploma, Mayo Clinic has opportunities for you.
View open positions
Remote
At Mayo Clinic, we offer a wide range of opportunities that can be done completely remote. We know there is no boundary when it comes to changing patient lives.
View open positionsTop-ranked in the U.S.
Mayo Clinic is top-ranked in more specialties than any other hospital according to U.S. News & World Report 2023-2024. More than 1.4 million patients from 140 countries are seen at Mayo Clinic locations each year.
Diversity, Equity & Inclusion
We lean on the variety of our colleagues’ perspectives and backgrounds to continuously challenge ourselves and to create a workplace that supports diversity, equity and inclusion. Become part of the legacy that embraces our differences and enables us to provide the best care to patients from all over the world.
Read MoreBenefits
As your career evolves, our compensation and benefits packages are designed to change with you—meeting needs now, and anticipating what comes next. We know that when Mayo Clinic takes care of you, you can take better care of our patients.
Read MoreLearning Opportunities
We offer a variety of opportunities to continue to learn and grow with Mayo Clinic. Ranging from internships to fellowship programs, there is no stopping where you could go next.
Read MoreJoin our talent community.
Join our global talent community to receive alerts when new life-changing opportunities become available.
Walk a path that can show you the world.
Search global careers
-

About Us
If you want to know what it's really like at Mayo Clinic, just ask. You'll find that our pride–in where we work, and in what we do–is a common trait. You will also find a lot of inspiring stories about lives changed for the better.
-
National Surgical Assistant Week
The Nurse Residency Program (NRP) is for all nurses with less than 12 months of experience, to be completed within the first year. NRP provides a framework for a successful transtion to a professional nurse by promoting educational and personal advancement.
-

Benefits
As your career evolves, our compensation and benefits packages are designed to change with you — meeting needs now, and anticipating what comes next. We know that when Mayo Clinic takes care of you, you can take better care of our patients.
Events
We would love to connect with you.
Click the button for a list of our upcoming events.
Equal opportunity
All qualified applicants will receive consideration for employment without regard to race, color, religion, sex, gender identity, sexual orientation, national origin, protected veteran status, or disability status. Learn more about "EEO is the Law." Mayo Clinic participates in E-Verify and may provide the Social Security Administration and, if necessary, the Department of Homeland Security with information from each new employee's Form I-9 to confirm work authorization.
Reasonable accommodations
Mayo Clinic provides reasonable accommodations to individuals with disabilities to increase opportunities and eliminate barriers to employment. If you need a reasonable accommodation in the application process; to access job postings, to apply for a job, for a job interview, for pre-employment testing, or with the onboarding process, please contact HR Connect at 507-266-0440 or 888-266-0440.
Job offers
Job offers are contingent upon successful completion of a post offer placement assessment including a urine drug screen, immunization review and tuberculin (TB) skin testing, if applicable.
Recruitment Fraud
Learn more about recruitment fraud and job scams
Advertising
Mayo Clinic is a not-for-profit organization and proceeds from Web advertising help support our mission. Mayo Clinic does not endorse any of the third party products and services advertised.
Advertising and sponsorship policy | Advertising and sponsorship opportunities
Reprint permissions
A single copy of these materials may be reprinted for noncommercial personal use only. "Mayo," "Mayo Clinic," "MayoClinic.org," "Mayo Clinic Healthy Living," and the triple-shield Mayo Clinic logo are trademarks of Mayo Foundation for Medical Education and Research.
Any use of this site constitutes your agreement to the Terms and Conditions and Privacy Policy linked below.
Terms and Conditions | Privacy Policy | Notice of Privacy Practices | Notice of Nondiscrimination
© 1998-2024 Mayo Foundation for Medical Education and Research (MFMER). All rights reserved.